The study aimed to evaluate the link between the IL-4-C590T polymorphism and asthma susceptibility in children by meta-analysis.
SourcesThe study collected all the case-control studies found in PubMed, Embase, CNKI, Wanfang, VIP, and other databases until September 2019. Stata v. 15.0 was used to conduct meta-analysis, calculate the combined OR and its 95% CI, and then conduct subgroup analysis.
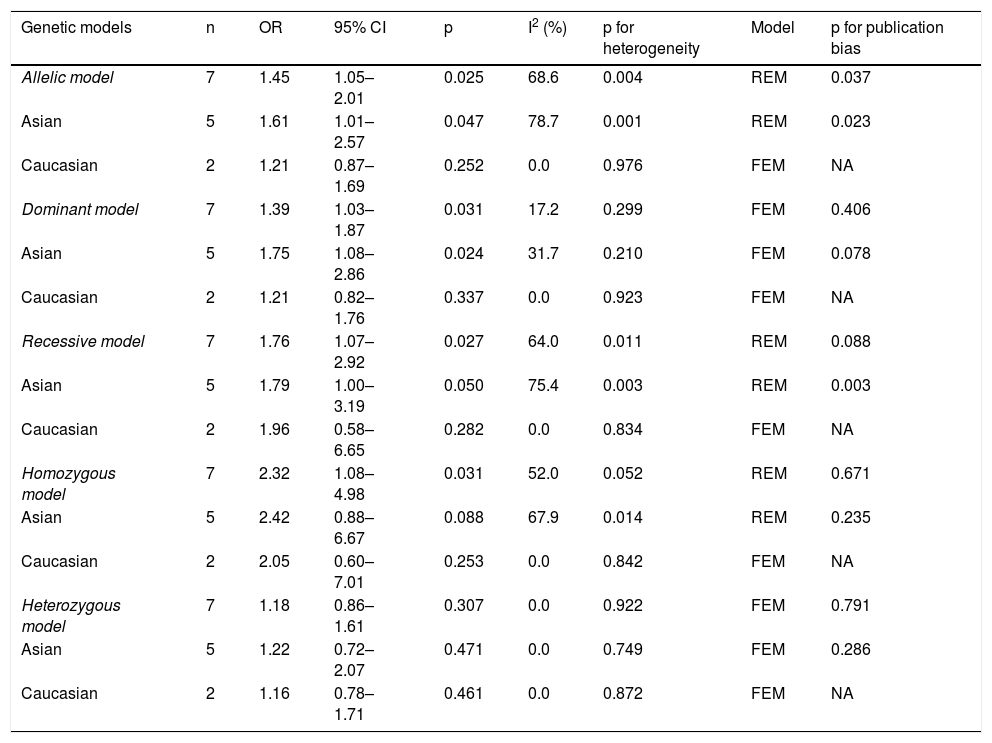
Summary of the findingsSeven studies were included in the study, containing 860 cases and 810 controls. Relative to the C allele, the T allele at the IL-4-C590T locus was associated with susceptibility to asthma in children (OR = 1.45, 95% CI: 1.05–2.01). The results of ethnicity subgroup analysis showed that there was statistical significance, with OR = 1.61 (95% CI: 1.01–2.57) in the Asian population. In the dominant and recessive genetic models, the overall test and the Asian population subgroup analysis were statistically significant. In the homozygous model, there was statistical significance, but no statistical significance in heterozygous model.
ConclusionsThe IL-4-C590T polymorphism was associated with asthma susceptibility, and T allele and TT genotype may increase the risk of asthma susceptibility in children, especially in the Asian population.
Bronchial asthma is a common chronic airway disease; the etiology is complex, involving various cells (such as eosinophils, mast cells, T lymphocytes, neutrophils, etc.) and a variety of cytokines, which are jointly involved in chronic airway inflammation.1,2 Asthma is an airway inflammatory alteration, which leads to airway structural reconstruction, airway space stenosis, airway hyper responsiveness (AHR), and airway obstruction in children with asthma.3,4 Studies have shown that5–7 the involvement of a variety of inflammatory factors and cytokines in the occurrence and development of bronchial asthma, the release of various types of inflammatory factors, and the infiltration of different airway inflammatory cells are the common pathological characteristics of the disease. The main clinical manifestations are paroxysmal dyspnea, wheezes, and diffuse wheezing. Asthma has genetic susceptibility, with multiple genes and environmental factors controlling its occurrence and development, as well as familial clustering and ethnic and geographical differences. Asthma can start at any age, but it is most common in early childhood. According to incomplete statistics, about 334 million people worldwide suffer from asthma. The overall prevalence rate of asthma in children under the age of 14 in China is 3.02%, and about 18,000 people die from asthma each year.8 Recurrent asthma affects the normal life of child patients, increases the family’s additional household financial expenditure, and wastes social resources. At present, most scholars believe that among the numerous factors causing asthma, gene may be the most vital factor, so it is a crucial task for researchers to study the pathogenesis and related susceptibility genes of bronchial asthma in children. Most scholars considered the functional imbalance of T helper type 1 and helper type 2 (Th1/Th2) cell subsets as the basic immunological alteration of asthma.9 Keeping the immune function of the human body in a normal condition is based on maintaining the Th1/Th2 cytokines in equilibrium. Once the balance is broken, there may be immune dysfunction and the occurrence of diseases (such as asthma). During the attack of asthma, the occurrence of autoimmunity can promote the differentiation of Th0 cells into Th2, resulting in hyper-function and breaking the balance between Th1 and Th2. Interleukin (IL) is a kind of cytokine which interacts between leukocytes or immune cells and has multiple functions, such as anti-inflammation and immunosuppression.10,11 It is also an important cytokine that affects children with asthma and IL-4 is the most representative of the IL family. IL-4 can promote the transformation of Th0 into Th2 cytokines and inhibit Th1, and facilitate the increase of the expression of immunoglobulin E (IgE) in children by enhancing the transformation and proliferation of B lymphocytes. The increase of humoral immunity of IgE and the lethality of natural killer (NK) cells further improve the chemotaxis and infiltration of eosinophils.12 Meanwhile, IL-4 can also react upon Th2 cells to increase the release quantity of IL-4, forming the malignant reaction to further aggravate the inflammatory response of children. Therefore, the study of IL-4 is a crucial point to explore in the pathogenesis of asthma. The base C of the IL-4 gene at the upstream 590 bp of ATG was mutated into base T, with three genotypes emerging: CC, CT, and TT, and the mutated allele T was positively correlated with the level of IgE. This means the change of the locus from C to T can cause an increase of the total serum IgE concentration; meanwhile, the increase of the total plasma IgE concentration is the main immunological characteristic of asthma. In the meantime, this study confirmed that the change of this locus of IL-4 from C to T could increase the concentration of total serum IgE; therefore, it was inferred that the single nucleotide variant of C-590T in the promoter region of IL-4 was associated with asthma in children.13 So far, there have been plenty of studies on the relationship between IL-4 gene single nucleotide variant and asthma susceptibility in children, both domestically and overseas. However, different scholars choose different gene loci; thus, the conclusions of the association are not consistent. In light of this, the C-590T locus of IL-4 cytokine was selected as the research object to explore the relationship between the C-590T locus and asthma susceptibility in children, in order to provide ideas for the study of the specific mechanism of asthma disease, as well as a theoretical foundation for the prevention and treatment of the disease.
MethodsLiterature retrievalThe study collected all the case-control studies that met the inclusion criteria through searching PubMed, the Excerpta Medica Database (Embase), the China National Knowledge Infrastructure (CNKI), Wanfang, the Database of Chinese Sci-tech Journals (VIP), and other databases until September of 2019. The keywords used in the search were IL-4, interleukin-4, single nucleotide variant, rs2243250, SNP, and asthma. The literature search was limited to English and Chinese, and the unpublished papers were excluded.
Inclusion and exclusion criteriaInclusion criteria: (1) the material should study the association between IL-4-C590T gene single nucleotide variant and asthma susceptibility in children; (2) the subjects in case-control studies were children; (3) the literature can provide data integrity or statistical indicators, OR, and 95% CI directly or indirectly; (4) all the literature in Chinese was published in the journals that are included in the core journals of the Peking University Library.
Exclusion criteria: (1) literature such as simple case study, case report, review, and evaluation; (2) redundant publications and literature with incomplete information; (3) studies with a control group that did not meet the Hardy–Weinberg equilibrium (HWE) were excluded, for there are several studies showing limitations about the HWE in several situations; (4) elimination of literature with a Newcastle–Ottawa scale (NOS) score of less than 6.
Data extractionData was independently screened and extracted by two researchers, and disagreements were solved by discussion or assistance from the third researcher. The contents of extraction included the following: first author, year of publication, country of study population, ethnicity, sample size of case group and control group, age of case group, numbers of genotype, genotyping method, control source, etc. Regarding the HWE of the control group, p < 0.05 showed that it was not in accordance with the HWE.
Literature quality evaluationResearchers read the full text of the literature carefully, and evaluated the quality of the literature according to the NOS.14 A score of less than 6 stars represents low-quality literature, and a score of 6 stars or greater represents high-quality literature. Only literature with a score of 6 stars or greater was included in this study. The two evaluators independently evaluated according to the uniform quality standards, extracted the information from literature, and then cross-checked. If there was any difference, it was resolved through discussion or with the assistance of the third researcher.
Statistical methodsThe data were collected according to the designed table to analyze the differences of alleles in the allele model between children with asthma and control children, and to analyze the differences of genotypes in dominant gene model, recessive gene model, homozygous genetic model, and heterozygous genetic model. Meta-analysis was carried out by using Stata v. 15.0 statistical software. Q-test was used to test the heterogeneity of the results. If I2 ≥ 50%, or p ≤ 0.05, indicating heterogeneity, the random effect model (REM) was used. If I2 < 50%, and p > 0.05, there was no heterogeneity. The fixed effect model (FEM) was then used for data analysis. The Z-test was used to test the significance of the combined OR. Subgroup analysis of ethnicity was performed. The meta-analysis was included in the evaluation of publication bias by using funnel diagram to determine whether it was symmetrical. The funnel diagram produces a diagram according to its OR and the standard error of each study (OR). There could have been publication bias if the funnel diagram was asymmetric, whereupon Egger’s test was used to test the publication bias.
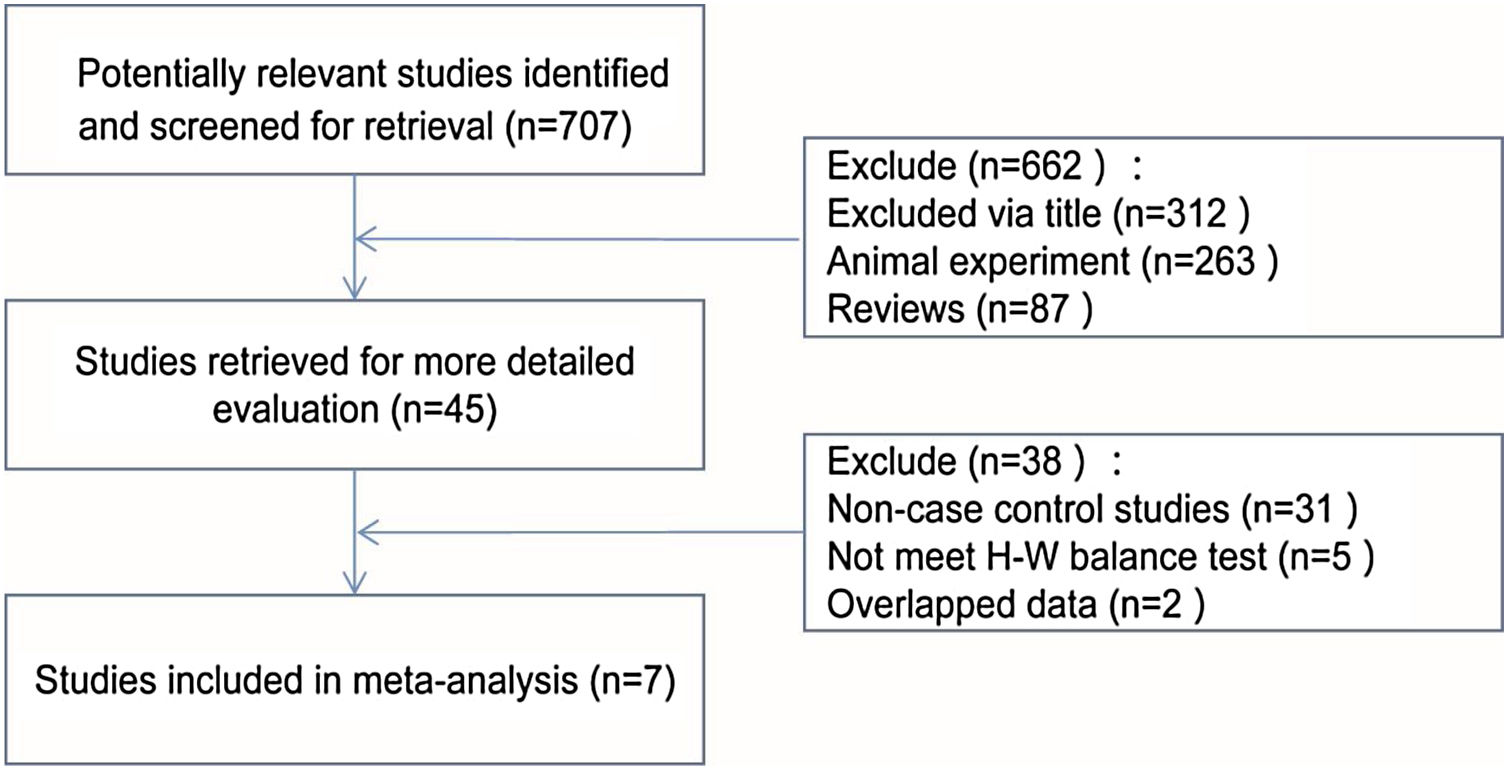
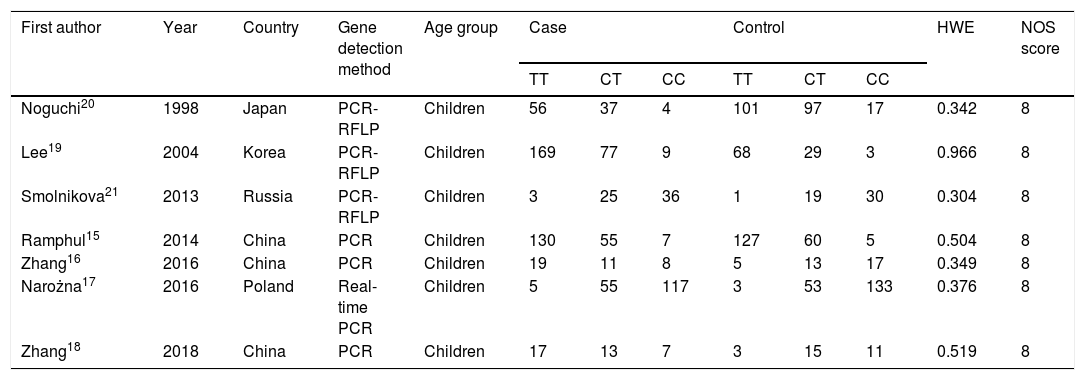
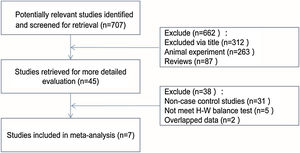
ResultsBasic information of research dataAccording to the criteria, a total of seven articles were selected,15–21 including two in Western countries and five in Asia, with a total of 860 patients in the asthma group and 810 patients in the control group. Details of the filter process were shown in Fig. 1. The characteristics of each study and the genotype distribution of the research reports is illustrated in Table 1.
Characteristics of the included studies.
| First author | Year | Country | Gene detection method | Age group | Case | Control | HWE | NOS score | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| TT | CT | CC | TT | CT | CC | |||||||
| Noguchi20 | 1998 | Japan | PCR-RFLP | Children | 56 | 37 | 4 | 101 | 97 | 17 | 0.342 | 8 |
| Lee19 | 2004 | Korea | PCR-RFLP | Children | 169 | 77 | 9 | 68 | 29 | 3 | 0.966 | 8 |
| Smolnikova21 | 2013 | Russia | PCR-RFLP | Children | 3 | 25 | 36 | 1 | 19 | 30 | 0.304 | 8 |
| Ramphul15 | 2014 | China | PCR | Children | 130 | 55 | 7 | 127 | 60 | 5 | 0.504 | 8 |
| Zhang16 | 2016 | China | PCR | Children | 19 | 11 | 8 | 5 | 13 | 17 | 0.349 | 8 |
| Narożna17 | 2016 | Poland | Real-time PCR | Children | 5 | 55 | 117 | 3 | 53 | 133 | 0.376 | 8 |
| Zhang18 | 2018 | China | PCR | Children | 17 | 13 | 7 | 3 | 15 | 11 | 0.519 | 8 |
PCR, polymerase chain reaction; RFLP, restriction fragment length polymorphism; HWE, Hardy–Weinburg equilibrium; NOS, Newcastle–Ottawa scale.
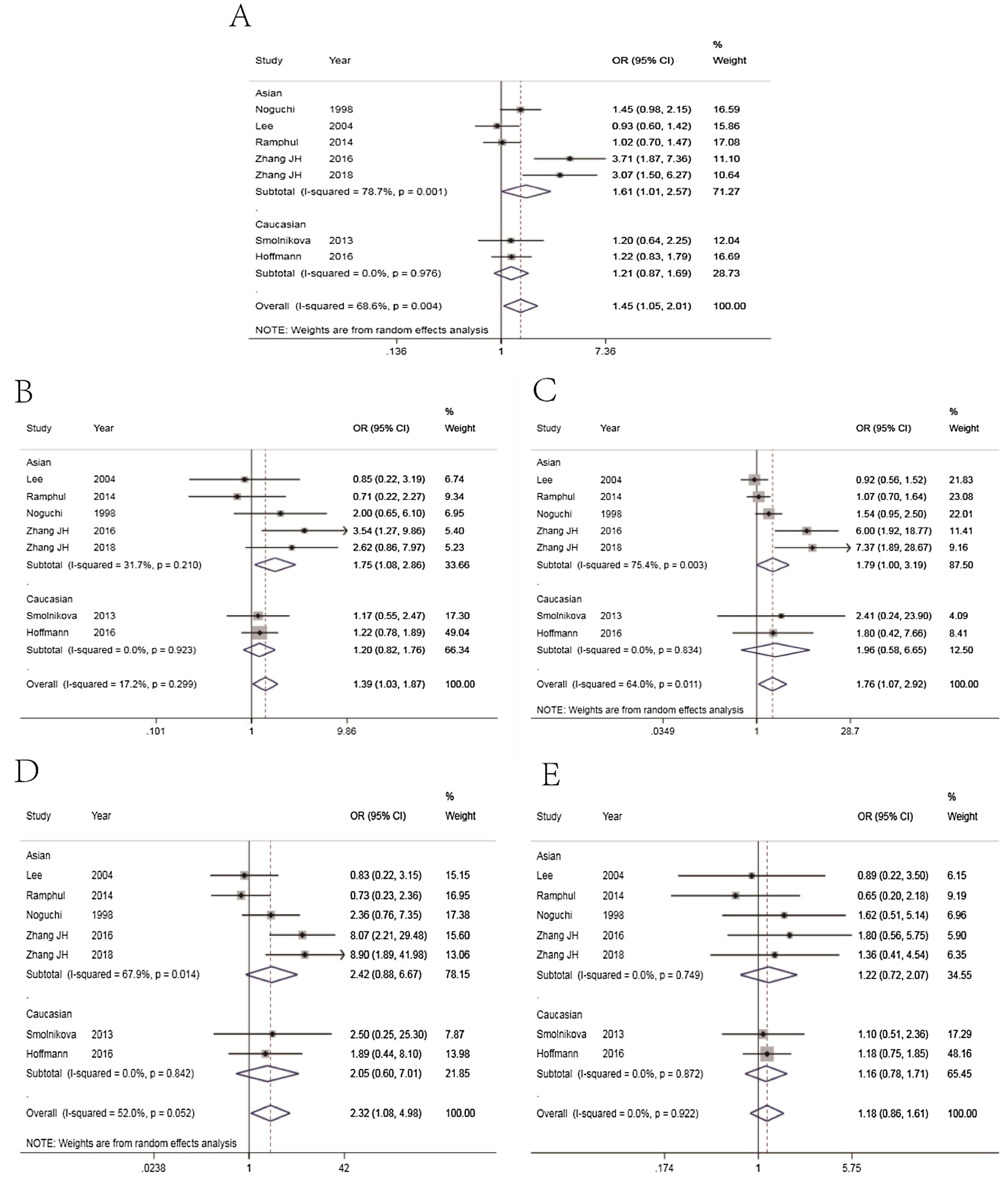
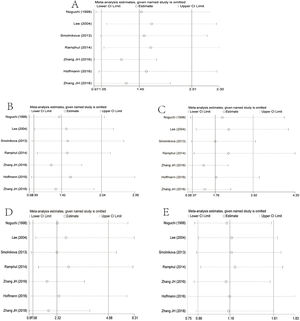
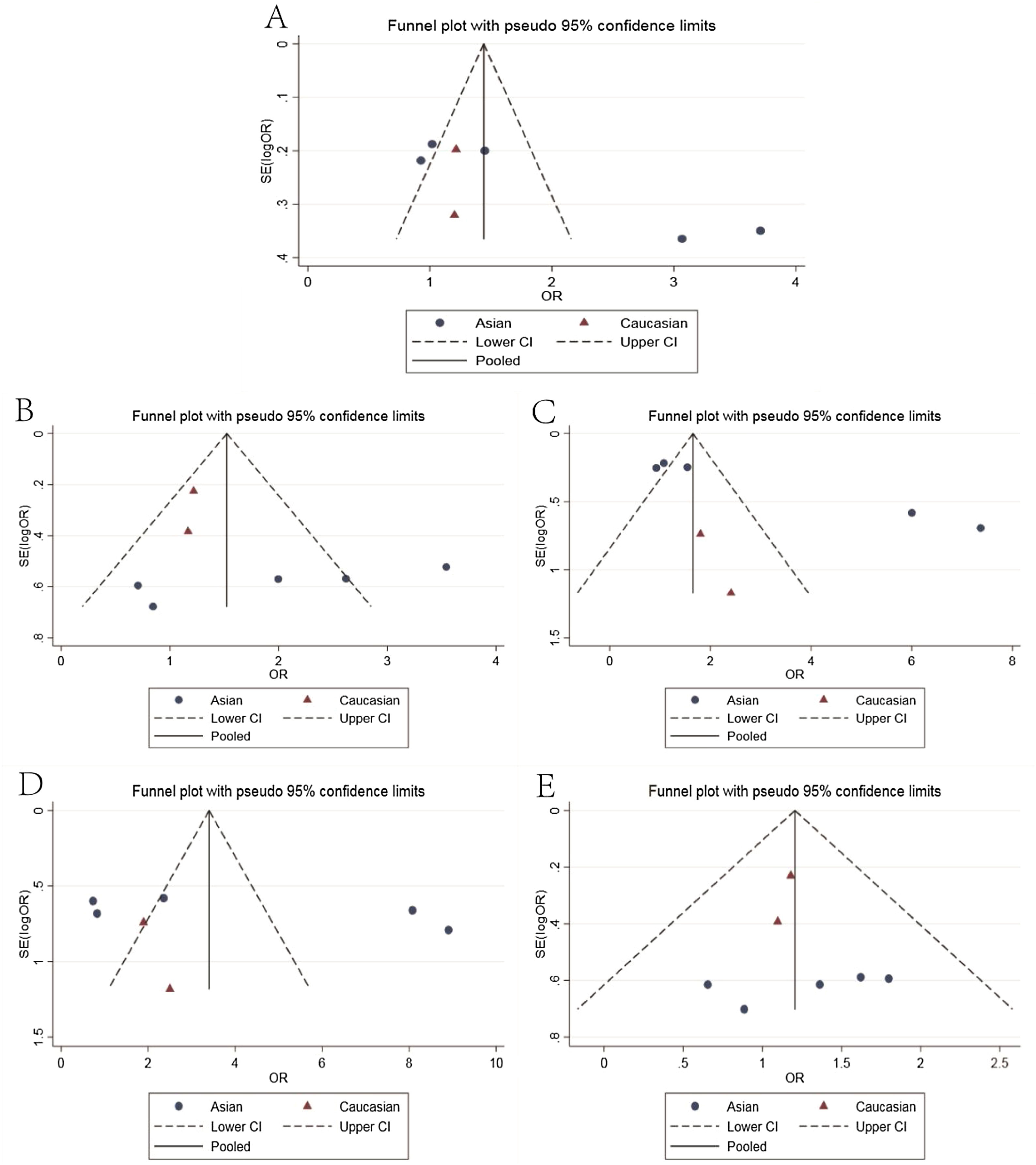
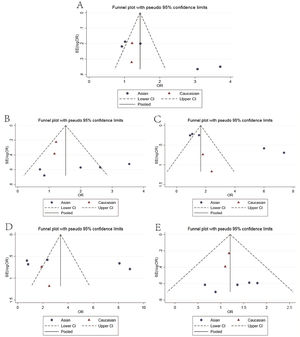
The main results of meta-analysis are demonstrated in Table 2 and Fig. 2. Relative to the C allele, the T allele showed I2 = 68.6%, (p < 0.05), indicating that there was significant heterogeneity among the studies, using the REM. The final results revealed that the combined OR = 1.45 (95% CI: 1.05–2.01, p < 0.05), indicating that the difference was statistically significant. According to ethnicity subgroup analysis, the results showed that the T allele was associated with susceptibility to asthma in children in the Caucasian population (OR = 1.21 95% CI: 0.87–1.69, p > 0.05) with statistically significant difference, and in the Asian population (OR = 1.61 95% CI: 1.01–2.57, p < 0.05) with statistically significant difference (Fig. 2A). When there was symmetrical deviation of funnel diagram (Fig. 3A), Egger’s test showed that there was partial publication bias (p < 0.05).
Results of meta-analysis for the IL-4-C-590 single nucleotide variant and childhood asthma.
| Genetic models | n | OR | 95% CI | p | I2 (%) | p for heterogeneity | Model | p for publication bias |
|---|---|---|---|---|---|---|---|---|
| Allelic model | 7 | 1.45 | 1.05–2.01 | 0.025 | 68.6 | 0.004 | REM | 0.037 |
| Asian | 5 | 1.61 | 1.01–2.57 | 0.047 | 78.7 | 0.001 | REM | 0.023 |
| Caucasian | 2 | 1.21 | 0.87–1.69 | 0.252 | 0.0 | 0.976 | FEM | NA |
| Dominant model | 7 | 1.39 | 1.03–1.87 | 0.031 | 17.2 | 0.299 | FEM | 0.406 |
| Asian | 5 | 1.75 | 1.08–2.86 | 0.024 | 31.7 | 0.210 | FEM | 0.078 |
| Caucasian | 2 | 1.21 | 0.82–1.76 | 0.337 | 0.0 | 0.923 | FEM | NA |
| Recessive model | 7 | 1.76 | 1.07–2.92 | 0.027 | 64.0 | 0.011 | REM | 0.088 |
| Asian | 5 | 1.79 | 1.00–3.19 | 0.050 | 75.4 | 0.003 | REM | 0.003 |
| Caucasian | 2 | 1.96 | 0.58–6.65 | 0.282 | 0.0 | 0.834 | FEM | NA |
| Homozygous model | 7 | 2.32 | 1.08–4.98 | 0.031 | 52.0 | 0.052 | REM | 0.671 |
| Asian | 5 | 2.42 | 0.88–6.67 | 0.088 | 67.9 | 0.014 | REM | 0.235 |
| Caucasian | 2 | 2.05 | 0.60–7.01 | 0.253 | 0.0 | 0.842 | FEM | NA |
| Heterozygous model | 7 | 1.18 | 0.86–1.61 | 0.307 | 0.0 | 0.922 | FEM | 0.791 |
| Asian | 5 | 1.22 | 0.72–2.07 | 0.471 | 0.0 | 0.749 | FEM | 0.286 |
| Caucasian | 2 | 1.16 | 0.78–1.71 | 0.461 | 0.0 | 0.872 | FEM | NA |
OR, odds ratio; REM, random effects model; FEM, fixed effects model.
The genotype TT + TC of the dominant genetic model relative to that of the genotype CC demonstrated I2 = 17.2% (p > 0.05), indicating that there was no statistically significant heterogeneity among the studies, and the FEM was used. The results showed that the difference was statistically significant (OR = 1.39, 95% CI: 1.03–1.87, p > 0.05). The analysis of ethnicity subgroup revealed that in Caucasians, there was no significant difference (OR = 1.21 95% CI: 0.82–1.76, p > 0.05); however, in the Asian population, the difference was statistically significant (OR = 1.75, 95% CI: 1.08–2.86, p < 0.05). The forest plot (Fig. 2B) suggests that the IL-4-C590 T gene single nucleotide variant was associated with asthma susceptibility in children, and that the TT + TC genotype raises the risk of asthma in children. The funnel plot was basically symmetrical (Fig. 3B). The Egger’s test results showed that the p-value was greater than 0.05, indicating that the publication bias was relatively low.
Recessive genetic modelsRelative to that of the TC + CC genotype, the TT genotype of the recessive genetic model demonstrated I2 = 64.0% (p < 0.05), indicating that the heterogeneity among the studies was statistically significant, adopting the REM. The results showed that there was significant difference (OR = 1.76, 95% CI: 1.07–2.92, p < 0.05). The analysis of ethnicity subgroups revealed that in Caucasians, OR = 1.96 was complicated with 95% CI: 0.586.65 and the difference was statistically significant (p > 0.05), and in the Asian population, the difference was statistically significant (OR = 1.79, 95% CI: 1.00–3.19, p < 0.05). The forest plot (Fig. 2C) demonstrated that the IL-4-C590T gene single nucleotide variant was related with asthma susceptibility in children, and the TT genotype increased the risk of asthma in children. The funnel diagram was basically symmetrical (Fig. 3C), Egger’s test shows p > 0.05, which indicated that the publication bias was well controlled.
Homozygous modelThe genotype TT, relative to CC, suggested that there was no statistically significant heterogeneity among the studies (I2 = 52.0%), and the REM was used. The final results showed that the difference was statistically significant (OR = 2.32, 95% CI: 1.08–4.98, p < 0.05). The analysis of ethnicity subgroups revealed that there was no significant difference in the Caucasian population (OR = 2.05, 95% CI: 0.60–7.01, p > 0.05) and similarly, in the Asian population (OR = 2.42, 95% CI: 0.88–6.67, p > 0.05). The forest plot (Fig. 2D) suggests that the IL-4-C590T gene single nucleotide variant was linked with asthma susceptibility in children, and that the TT genotype, compared with CC genotype, raises the risk of asthma in children. The funnel diagram was basically symmetrical (Fig. 3D). Egger’s test revealed that the publication bias was well controlled (p > 0.05).
Heterozygous modelCompared with CC, the TC genotype indicated that there was no statistically significant heterogeneity among the studies (I2 = 0%, p > 0.05), using the FEM. The results showed that no statistically significant difference was found (OR = 1.18 95% CI: 0.86–1.61, p > 0.05). The racial subgroup analysis revealed that there was no significant difference in the Caucasian population (OR = 1.16, 95% CI: 0.78–1.71, p > 0.05) and the same was found in the Asian population (OR = 1.22, 95% CI: 0.72–2.07, p > 0.05). The forest plot (Fig. 2E) implies that it cannot be concluded that the risk of asthma in children with the TC genotype was different from that of the CC genotype. The funnel plot was basically symmetrical (Fig. 3E). The Egger’s Test results suggested that there was no statistical significance, implying that the publication bias was relatively well controlled.
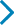
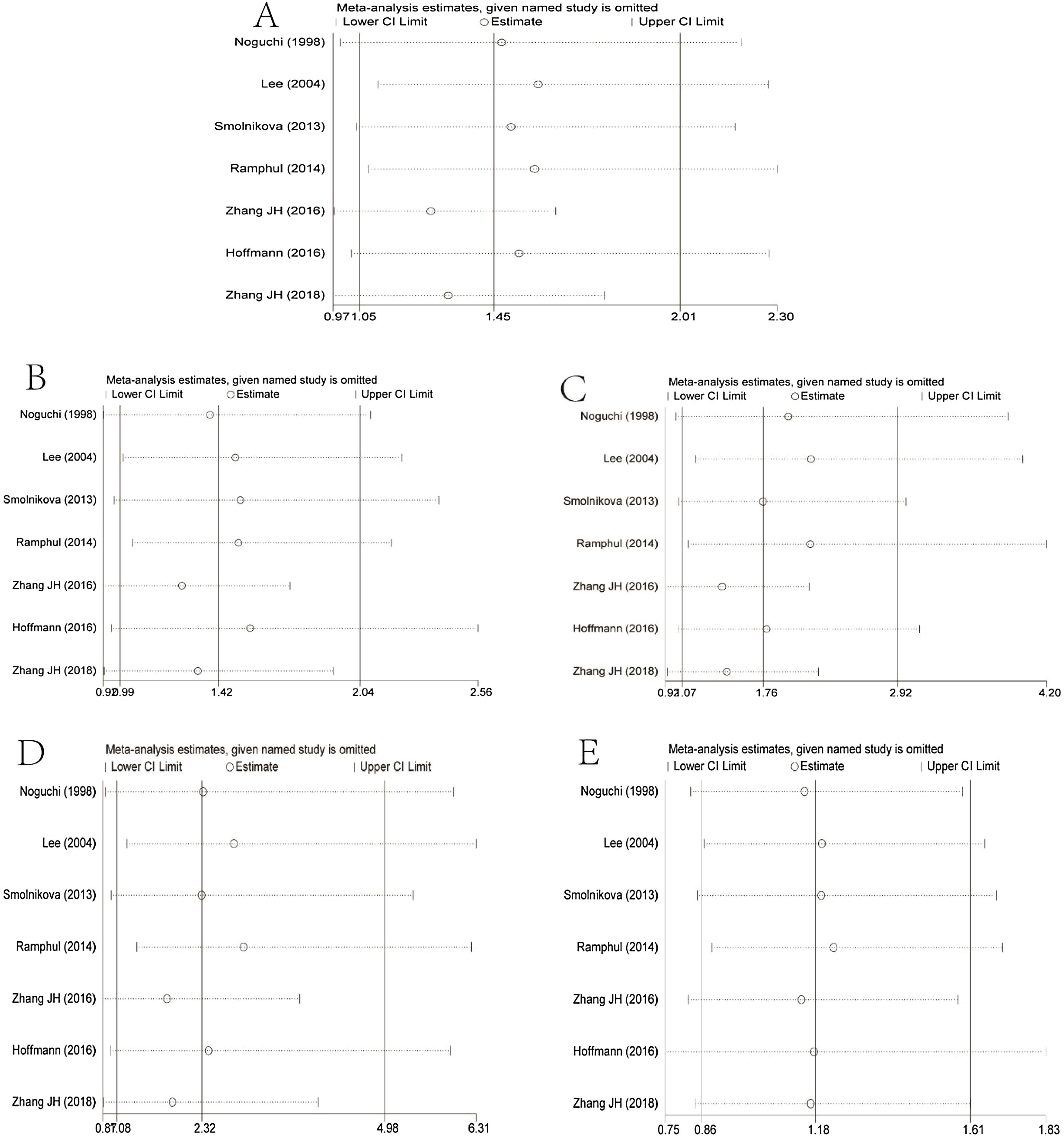
Sensitivity analysisFor the results of sensitivity analysis, see Fig. 4. The authors excluded each study one-by-one and used meta-analysis to explore the impact of a single study on the overall results. The results showed that there were statistical differences in the allele model, dominant gene inheritance model, recessive gene inheritance model, and homozygous inheritance model. The difference of the heterozygous genetic model did not change after being excluded from a single study. This shows that the conclusion is unstable to a certain extent.
DiscussionAsthma is a complex immune disease in which numerous cytokines and inflammatory mediators are involved in the pathogenesis. The inflammatory reaction is the pathophysiological basis of airway injury in patients with bronchial asthma, mainly due to the infiltration of eosinophils, mast cells, and T cells, as well as the effect of cytokines. Long-term chronic inflammation and damaging and rebuilding of airway epithelium, including changes such as increased airway smooth muscle, hyperplasia of goblet cells and fibroblasts, and subepithelial fibrosis play a significant role in the occurrence, development, and late irreversible alterations of asthma.22 As a common and frequently-occurring disease of the respiratory tract, it has attracted widespread attention from scholars because of its complicated etiology, recurrence, and the increasing incidence rate year-by-year in developing countries. Asthma is one of the recognized challenges in the world, and its pathogenesis is still unclear, but recent studies have shown that the imbalance between T cells (Thl/Th2) eventually leads to the occurrence of asthma.23–25 Immunology demonstrates that under normal circumstances, the Thl/Th2 in the body is in a stable state, but under the influence/effects of some external factors, the immune system will be undermined, resulting in shifting from Th1 to Th2. This increases the amount of Th2 cytokines (IL-4, IL-5, IL-13) and others. As these factors increase, they stimulate B lymphocytes in the body, resulting in increased IgE production, and EOS activation and proliferation in the meantime, as well as releasing various pro-inflammatory mediators, etc., causing anaphylaxis and damaging autologous tissues and functions. Th1 and Th2 cells in normal humans are differentiated from Th0 cells. The variety and direction of their differentiation is influenced by several factors. IL-4 plays a leading role in the transformation of Th0 to Th2 cells in the in vivo environment. Related studies have shown that the transformation direction of Th0 cells is also affected by other factors, such as local factors in the tissues, immunoreactive hormones, types and concentrations of antigens in the body, and types of antigen-presenting cells.26,27 In addition, the occurrence of asthma is related to other cells, such as cell surface molecules, inflammatory mediators, immunocompetent cells, and inflammatory cells.28 In recent years, with the development of the economy and the constant progress in human genetics research, studies on asthma-related genes have also continuously developed. Asthma is a polygenic genetic disease. The interaction between the environment and genetic factors is closely related to the occurrence of asthma, which is a common chronic respiratory disease in childhood,29 In order to provide proof for evidence-based medicine for clinical guidance, this study conducted a comprehensive analysis of the relationship between the IL-4-C590 single nucleotide variant and susceptibility to asthma in children.
A total of seven articles15–21 were included in the study, of which five were from Asian studies and two were from European and American studies. The results showed a correlation between the IL-4-C590T gene polymorphism and childhood asthma susceptibility. The T allele at the IL-4-C590T locus vs. the C allele was correlated with susceptibility to asthma in children. It is also statistically significant in the Asian population. In the dominant and recessive genetic models, the overall test and the Asian population subgroup analysis were statistically significant. In the homozygous model, the total comparative difference was statistically significant, but the subgroup analysis was not statistically significant. There was no statistical significance in the heterozygous genetic model. The results of publication bias showed that the publication bias of each genetic model was relatively small. The results of the heterogeneity test showed that I2 was typically less than 50%, indicating that the heterogeneity between the studies was small. Sensitivity analysis of genetic models of each gene showed that except for the heterozygous genetic model, there were certain instabilities that existed in the allele models, and the dominant, recessive, and homozygous genetic models. Ren et al.30 and other studies on the IL-4-C590T locus single nucleotide variant and susceptibility to asthma in children suggest that all five genotypes can increase the risk of bronchial asthma in children, which is inconsistent with the results of this study. One reason could be that this study incorporates the latest high-quality literature, and the second reason may be the exclusion of literature that did not meet the HWE test. It can be considered that there is association between the IL-4-C590T locus single nucleotide variant and asthma susceptibility in children with increased risk of asthma susceptibility in patients. However, considering the results of the sensitivity analysis, this conclusion should be cautious.
The study also had certain limitations: (1) The studies included were still relatively limited, mainly in Asia, Europe, and the United States. The number of studies included was also limited. There were no studies that focused on populations such as that of Africa. (2) In addition to the heterozygous genetic model, certain instabilities existed in the sensitivity analyses of other genetic models. (3) The linkage between genes and the interaction of genes and the environment on childhood asthma has not been analyzed. (4) In addition, given the number of studies included in this meta-analysis, researchers will need to wait more time to obtain more information for analysis in the future.
ConclusionsIn conclusion, the IL-4-C590T gene single nucleotide variant was associated with childhood asthma susceptibility, the T allele was associated with an increased risk relative to the C allele, and the TT and TT + TC genotypes were also susceptible genotypes, especially in Asians, but not in Caucasians. However, considering that there were still some limitations in the research, such as sensitivity issues, lack of considering the gene-gene interactions, genes and the environment, etc. Therefore, more in-depth research on a larger scale as follow-up is still needed.
Conflicts of interestThe authors declare no conflicts of interest.